There are many, many more insects in the world than are known to science. It is estimated that about 1 million insect species have been classified to date: that is, given valid scientific names, with voucher specimens hopefully kept for comparison in museums around the world. However, it has been estimated that there could be as many as 5.5 million insects (range: 2.6–7.8 million) in the world (Stork, 2018), so there are still a huge number to discover. A large proportion of these will probably be flies and beetles, which is very good thing in my opinion!
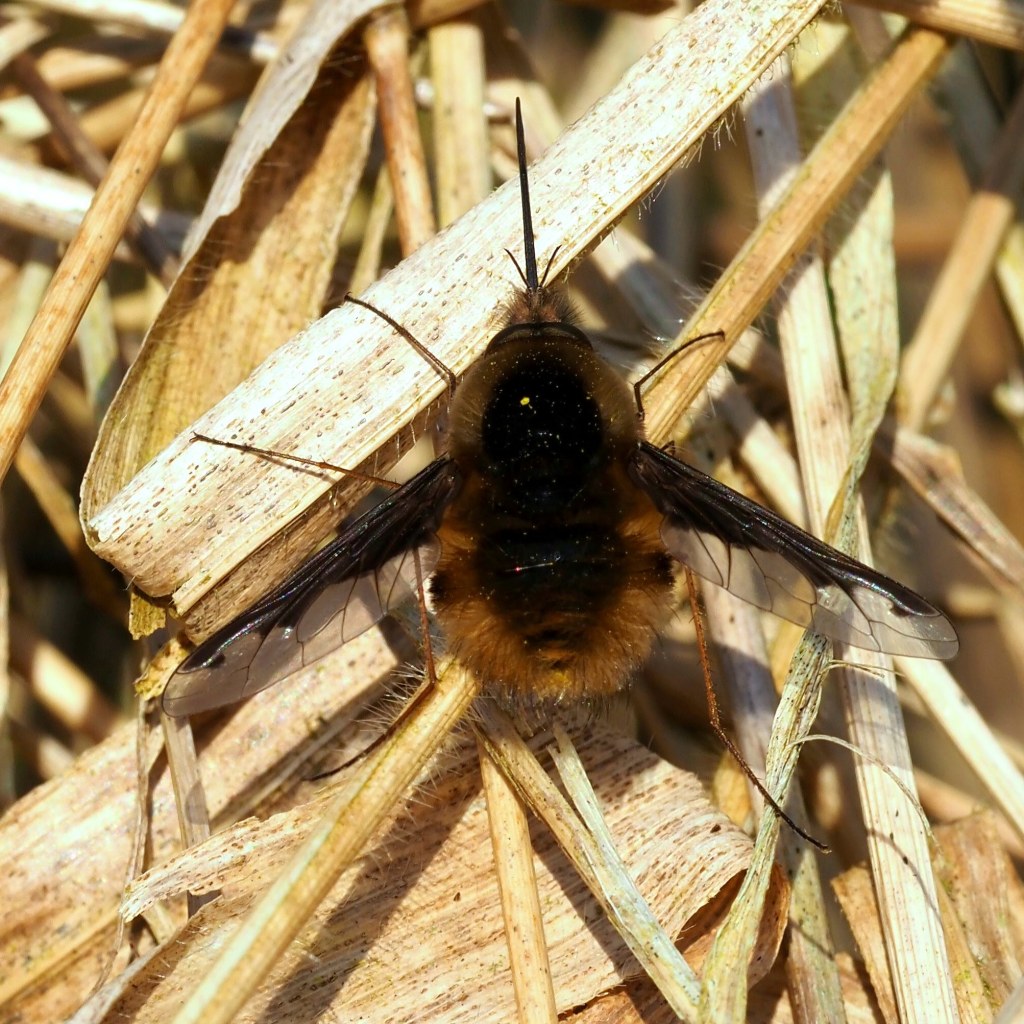
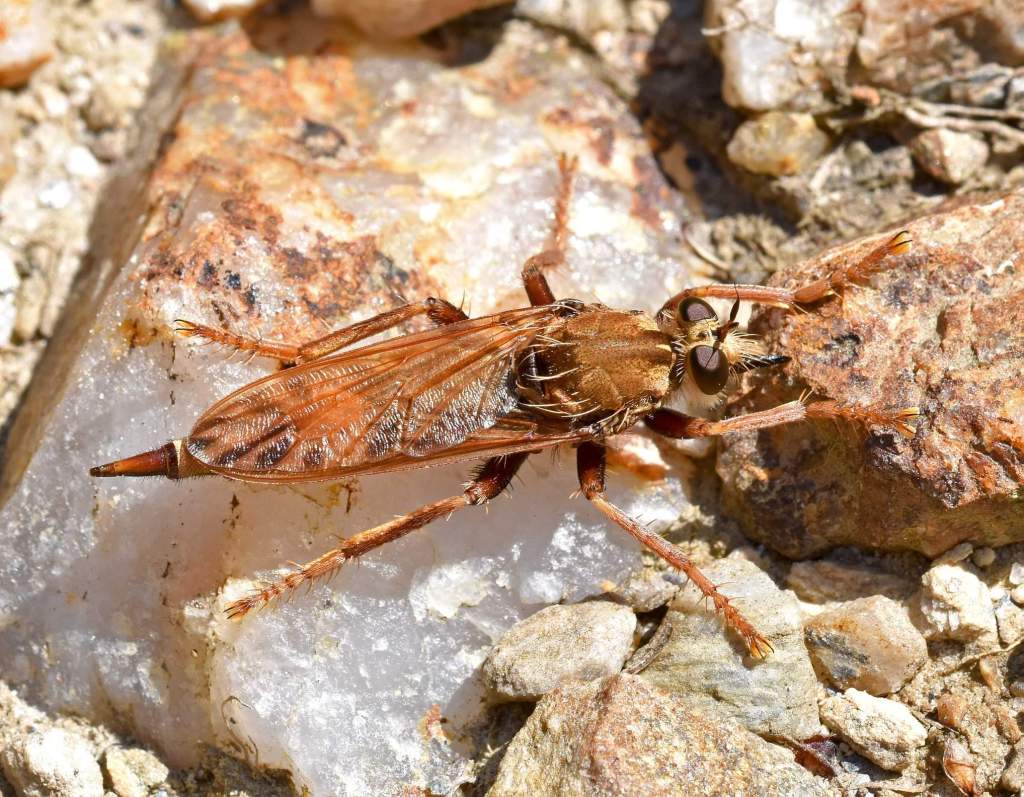
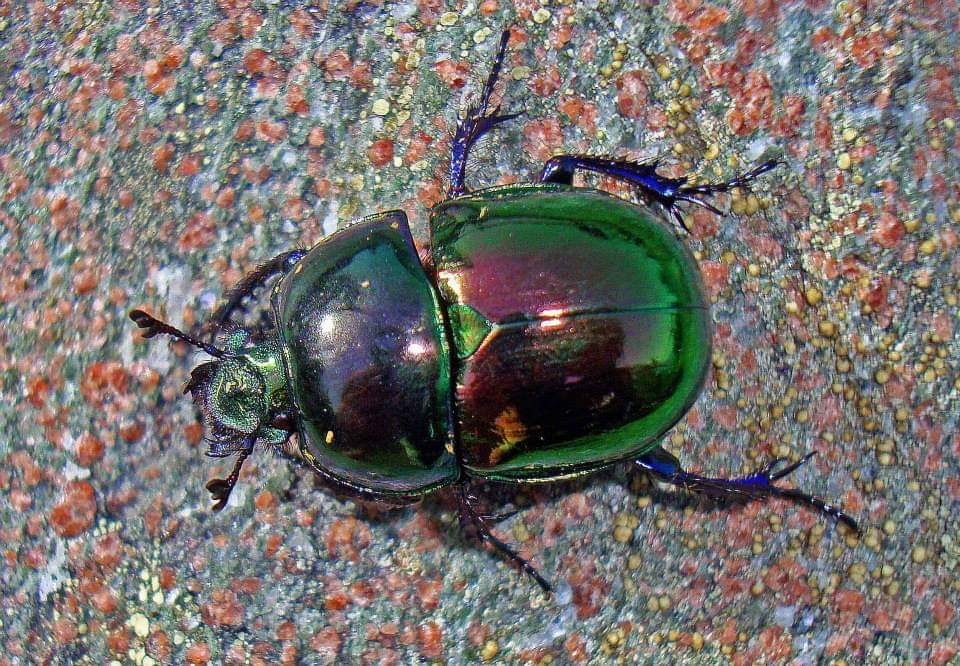
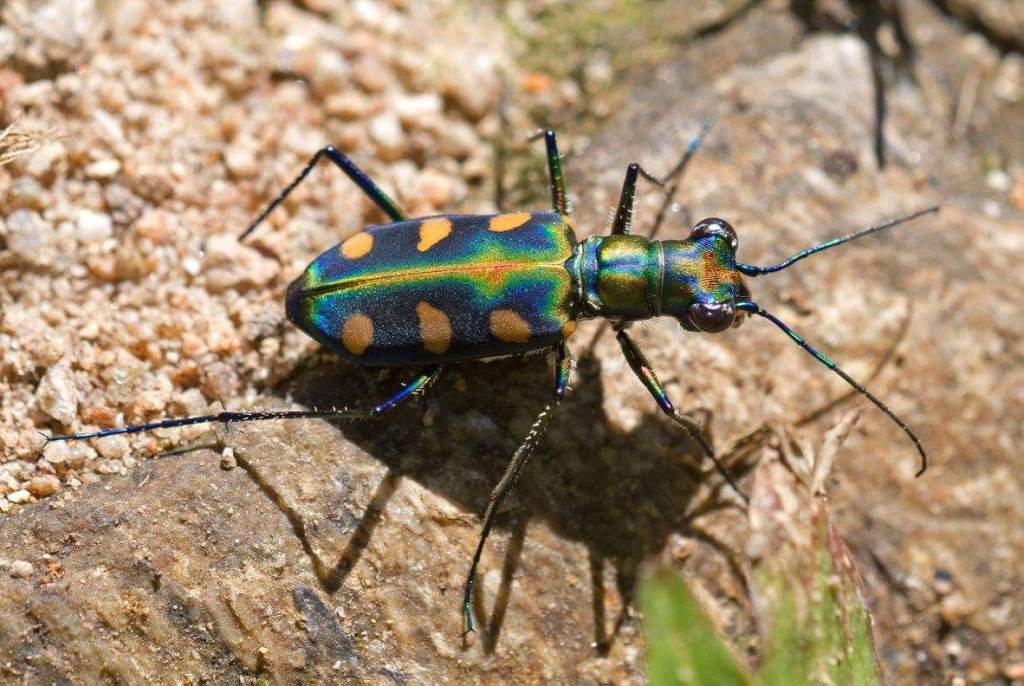




Although, thousands of new species are discovered and named each year, the rate of progress using traditional taxonomic methods is far too slow to examine and catalogue all of the – 80 to 90%? – remaining species within a reasonable time frame; and a large proportion of the currently unknown species are liable to disappear, along with their habitats, before they can be documented. Time is running out and there are simply too few specialist taxonomists to do the work; plus, there is also a worrying deficit of young taxonomists (Hochkirch et al., 2022). In other words, the availability of experts with the knowledge and capability of identifying and describing new insect species is declining, at the same time as the insects themselves are in danger of going extinct! But all is not lost, thanks to modern science, as newer, faster ways of distinguishing biodiversity are applied.
Some readers may be surprised to learn that biologists are divided about what constitues a species. The ‘species problem’, as it is called, is that there are a multiplicity of species concepts and no agreed concensus as to which one is adequate. For example, one taxonomist might use a species concept based on morphological features (i.e. physical structures like male genitalia, see below), while another may use genetic similarity based on comparisons of DNA sequences (described below), and yet another might use a concept based on phylogeny (Richards, 2013).
A simple definition of a species is a group of organisms that can reproduce with one another in nature and produce fertile offspring. However, with the advent and widespread application of molecular methods, such as DNA barcoding (see below), scientists can categorise species in a different way, and have been discovering many cryptic, or previously ‘hidden’ species. Cryptic species are one or more – often many more – identical, that is to say morphologically indistinguishable, biological groups that are genetically quite distinct and incapable of interbreeding. So, what were once thought to be single species, are now being discovered to be reproductively isolated species, sometimes called sibling species.
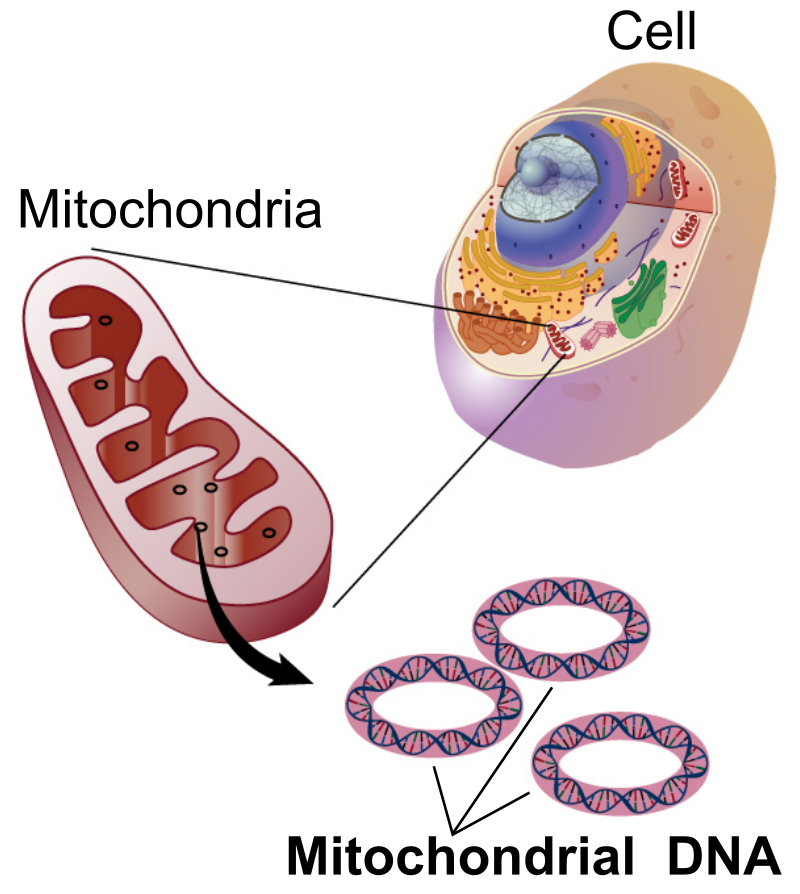
A region often used for DNA barcoding in insects is a segment of a mitochondrial gene (mtCOI) which contains the instructions to make a large protein, called cytochrome c oxidase. The mitochondrial genome of animals evolves more rapidly than genes within the nucleus of a cell, so the changes which accumulate in the mitochondrial DNA (see illustration below) are useful for studying patterns of variation among populations (Herbert et al., 2003).
Cryptic species may have been overlooked by taxonomists who first described a species, simply because there were no differences in the morphological features of the stage, or sex, of the insect, that they were examining (Chenuil et al., 2019). For example, both immature and female caddisflies (Trichoptera) are almost impossible to identify; they all look the same (Zhou et al., 2011). If taxonomists could collect all of the life stages of a given species, from throughout its geographical range, and examine all of the various anatomical features, they might be able to discern some different types, or perhaps some new species. However, even such enormous efforts like this may still fail to discern cryptic diversity, as it is called, if the species keep themseves reproductively isolated by means of their behaviours. For example, singing different calling or courtship songs (Cannon, 2023). I.e. behavioural characteristics that leave no discernable physical differences.
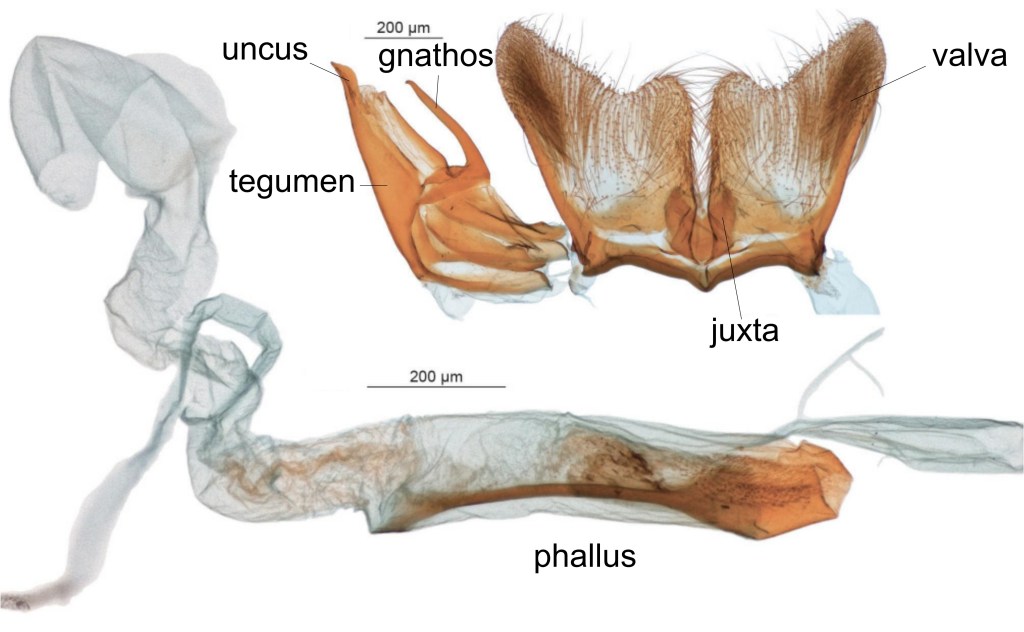
A nice of example of where cryptic butterfly species were discoved by examining female (rather than male) genitalia, is provided by Afrotropical Melanitini butterflies like Gnophodes (see below) and Melanitis (Pyrcz et al., 2020). According to these researchers, male genitalia vary little between both the species, and genera of Melanitini, but female genitalia provide more “consistent, informative characters, providing a firm basis for distinguishing between the genera, and between species that are externally quite similar” (Pyrcz et al., 2020).
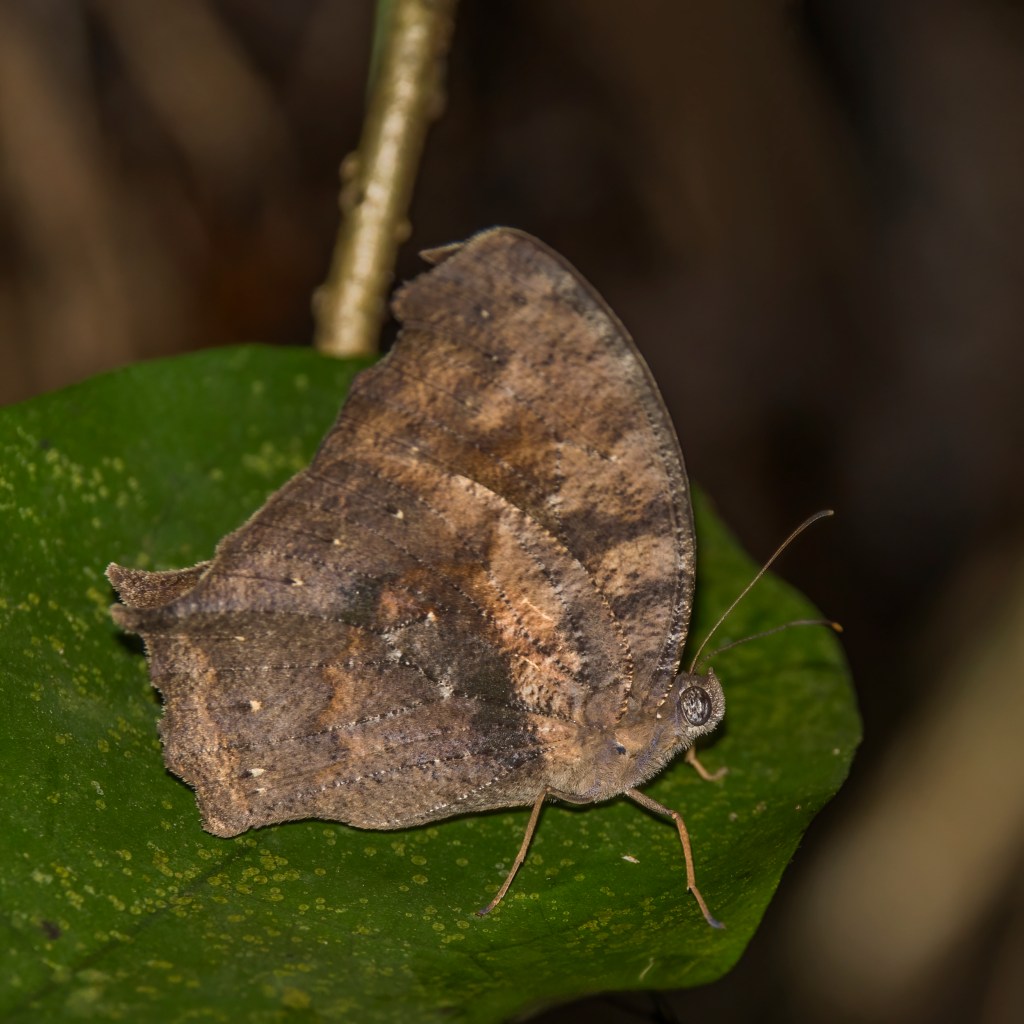

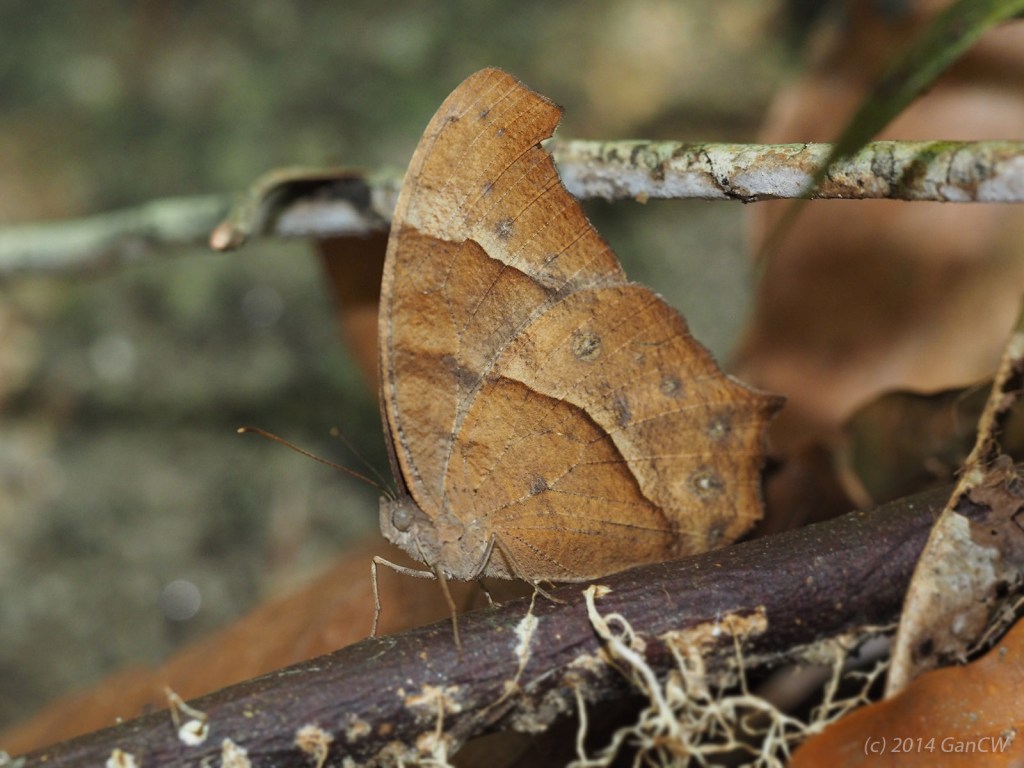
A combination of molecular (mitochondrial gene COI DNA) and morphological (using adults and larvae) methods revealed that this tribe of African butterflies, Melanitini, is much more diverse than was previously recognised, with four new genera and twice as many species (Pyrcz et al., 2020). Some species, like Melanitis leda, the common evening brown, have a very wide geographical distribution – including Africa, South Asia and South-east Asia extending to parts of Australia – with many different subspecies (although these are not necessarily cryptic species).
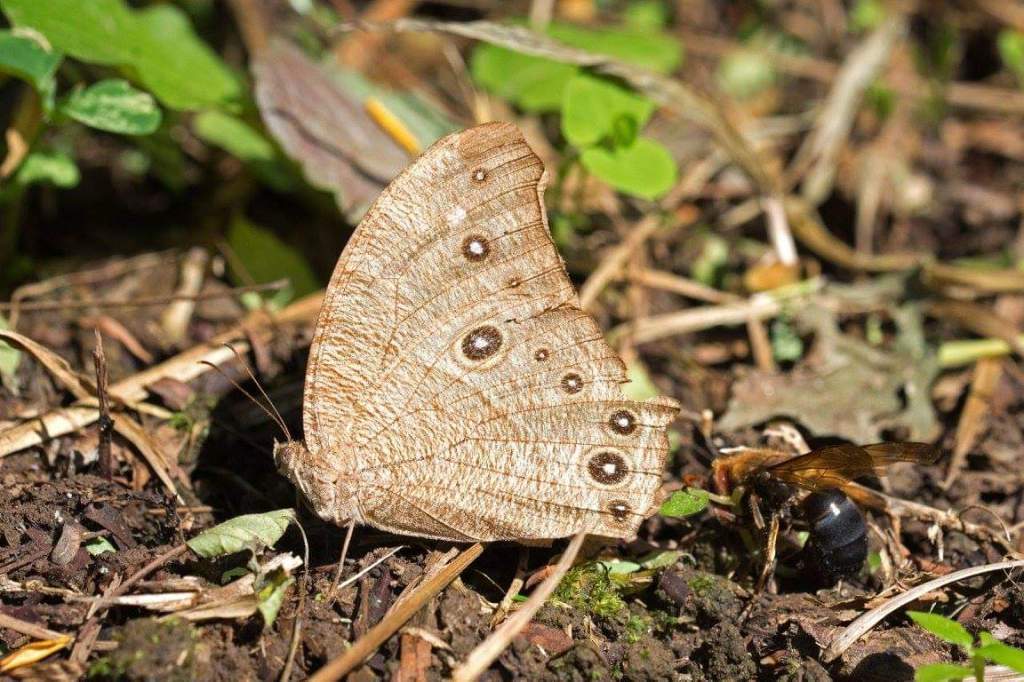

The dry season form (DSF) of this polymorphic butterfly (below), is very different in appearance from the wet season form (above), when comparing the undersides of the wings. The DSF of this butterfly lacks the ptominent eye-spots of the WSF, and relies much more on crypsis – blending in with dry leaves – for survival.

https://www.flickr.com/photos/gancw1/13004815614/in/photolist-kPc5AS-8hR3PK-2hZs1Q7-21a18kD-2hZs1RQ-2hZtaV5-21q8KGw-EnxFdf-Hyd4oZ-didykB-22sa2cf-EnxwGY-8FVsxt-6U3KmV-219ZYZF-788VXa-6e2hdR-6ecRVh-gdE6pz-6e7oZ5-AmtAnS-gdEtKW-P7E6S1-2mmXLbc-tiVcmA-9mPv2Q-dfnMLb-hvcKwS-2hYr1Az-9x8bEL-dV1cz7-2mjoQ3L-j7Gmrq-6Bg735-8hP4DT-6ebsc6-qt9fSz-L9cRb3-fTvrqg-eYFvSY-gfaHhJ-28uiyGb-6Bg6M3-2nVnvVR-dxwKFL-aiin3N-6BbVuT-gx7voX-6Bg6C5-dKXYi6
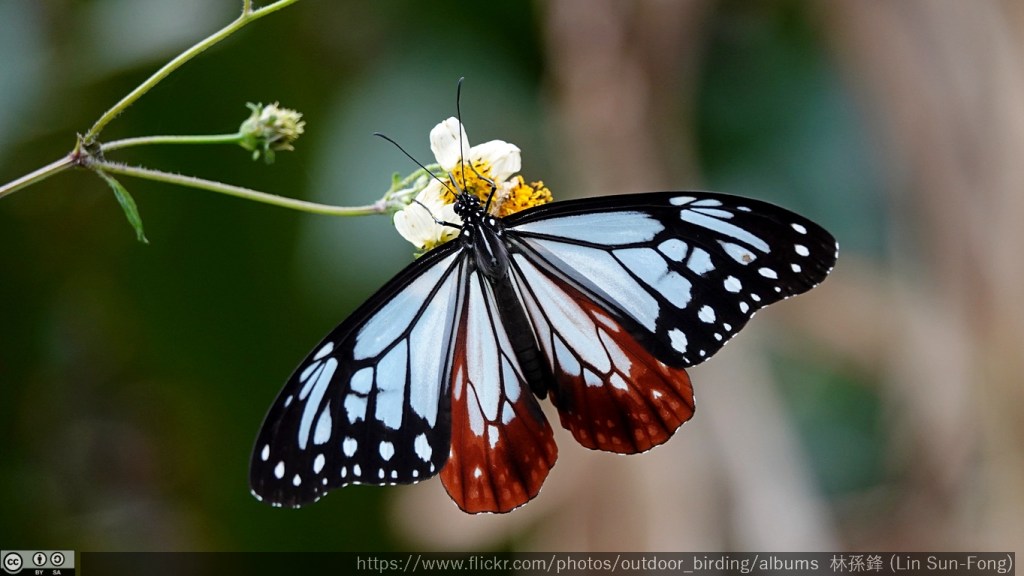
A recent estimate concluded that each morphology-based insect species contains (on average) 3.1 cryptic species (Li and Wiens, 2022). This implies that there may be three times as many extant species, on average, than was was estimated using non-molecular methods. However, it is likely that in many cases, once species have been separated by DNA barcoding, morphological differences between them may be found by further research, in adults or other life stages, such as larvae (Stork, 2018). One recent example of this cryptic diversity, is that of the chestnut tiger butterfly, Parantica sita (Kollar) (Nymphalidae, Danainae) in China. This species was recently split into three species, using molecular divergence and genitalia differences (Hu et al. 2022).

Lin Sun-Fong https://www.flickr.com/photos/outdoor_birding/51921343778/in/photolist-gMe1gi-gMd2iw-2mUshJG-2mRGjpu-2n243FV-2n292xB-2n2aDE2-zTRBBH-61qZik-yf6q2b-NgbADA-Cmj5Fv-2mURNDM-zkNLdh-2kXivoW-MVkbbG-MYn1NG-CEw4C3-5zrJKn-4hH4LU-4hH3Ru-H57yJb-fwzsEH-7ZtKCM-7ZtKNi-ax8f82-axag6F-axag6a-axcY2G-mXR36-axag7i-axag8D-2n78FqR-mXRtJ-7ZwVsw-7ZwVV7-4iAQXg-BssaA5-AX4WHo-auqV6J-aunuUz-euWJt-euWJs-euWpf-oPVFx-oPVFB-euWpe-oPVFF-oPVFD-2n77sAo
A large number of endangered species are probably cryptic species complexes, which effectively enhances their degree endangerment, since the cryptic species within the complex will be fewer in number, and have smaller distributions than the nominate species. For butterflies in genera, Janzen et al. (2028) ‘hazard’ an estimate that that ’10–20% of the traditional,
morphologically based “single” species will turn out to be two
or more’.
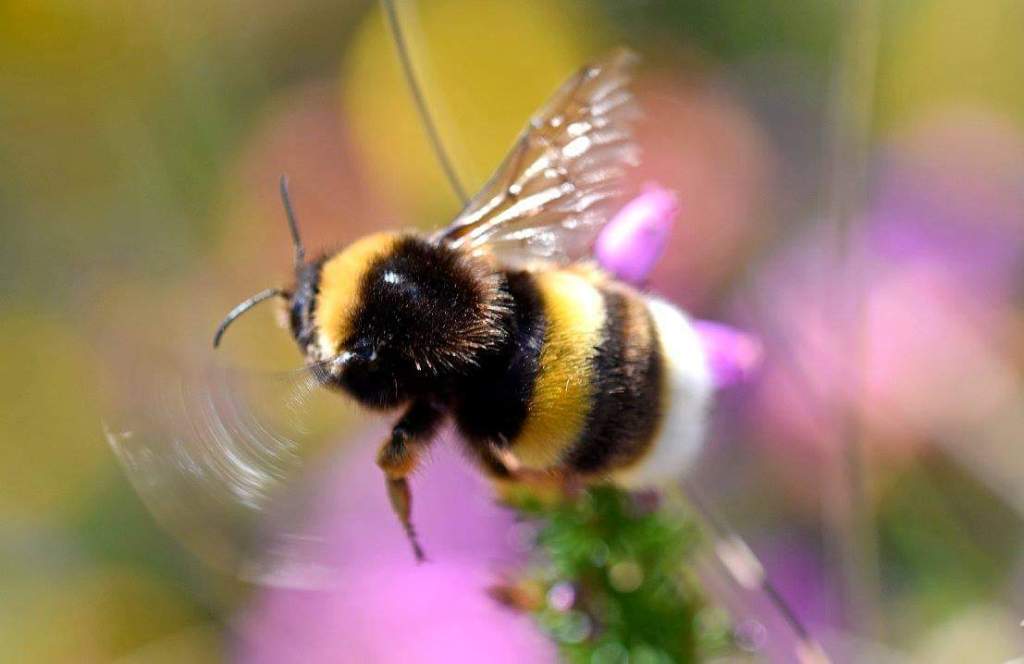
There are many examples of cryptic diversity in bumblebees. For example, the so-called, Bombus lucorum complex, contains three closely related species (Bombus lucorum, B. magnus, and B. cryptarum) that are very similar in colour, and in terms of their male genitalia (Waters et al., 2012). Their status as separate species has however, been confirmed by DNA-based identification methods (i.e. nucleotide sequence data), particularly the use of COI barcoding. There have been some suggestions that the queens of these three bumblebee species can be told apart based on the extent of the yellow collar on the thorax (Bossert, 2014), although other studies have contested this (Carolan et a., 2012). See illustration (below).
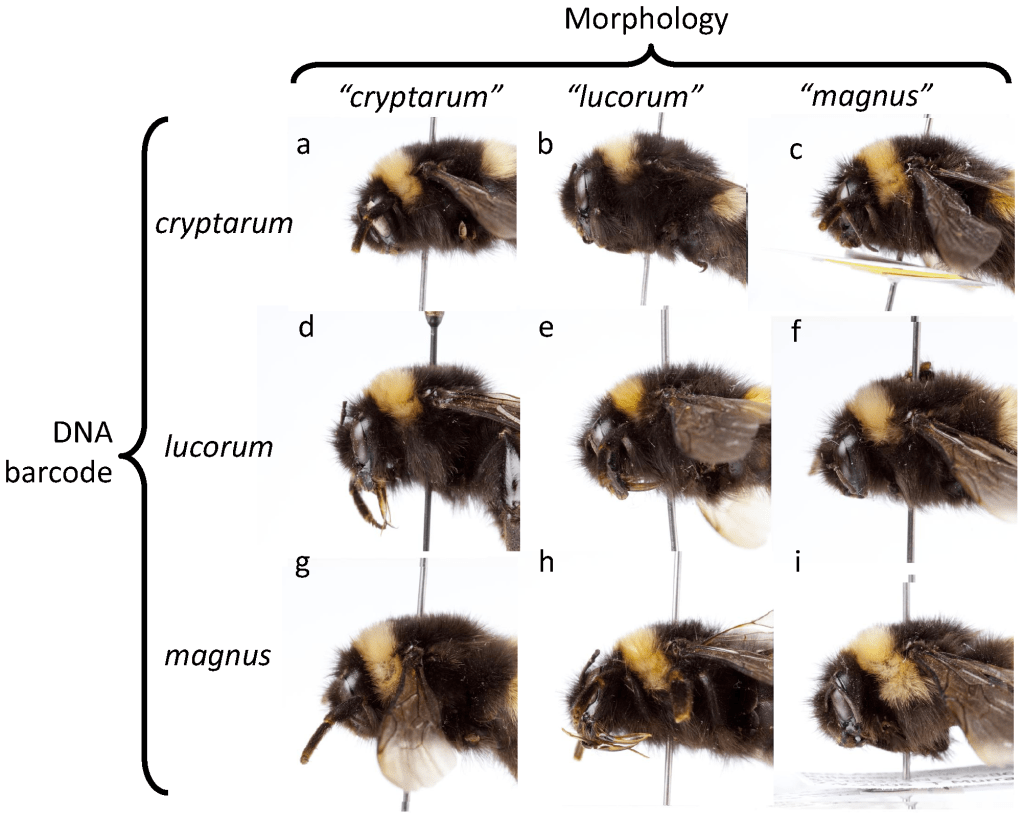
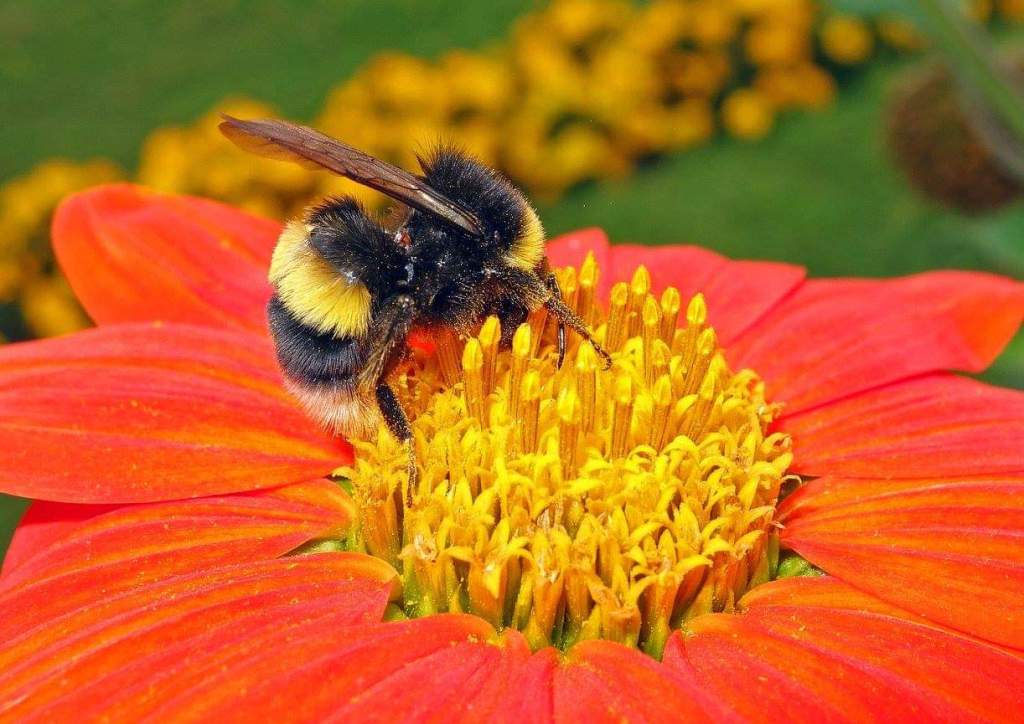
Bombus lucorum appears to be the most common and widespread of the three species, and is present throughout mainland Great Britain. B. magnus is a generally larger bee (at least the queens and males are larger; not the workers) and is strongly associated with heather, Calluna vulgaris. Thus, B. magnus and is of greater conservation concern than B. lucorum because of its restricted distribution and reliance on heather. I was so amazed at the size of a large, white-tailed bumblebee queen I encountered on the heather-clad heathlands of Galicia, Spain (see below), I felt certain that it was Bombus magnus!


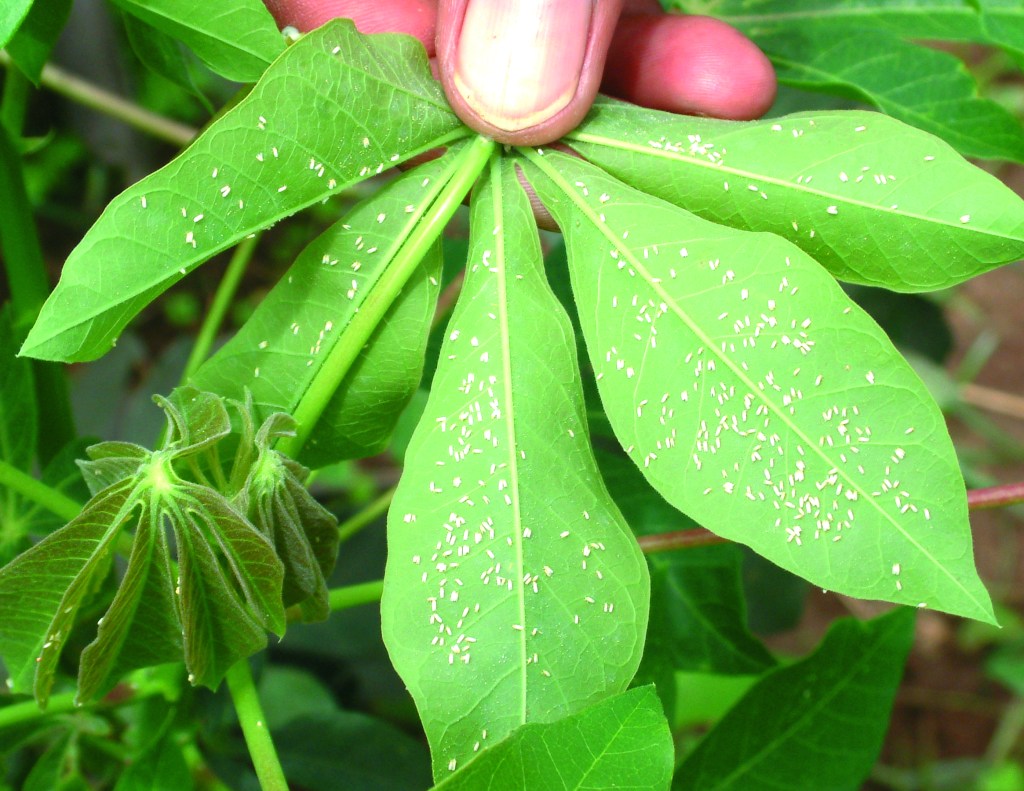
The whitefly, Bemisia tabaci Gennadius (Hemiptera: Aleyrodidae) is another species with marked cryptic diversity. A worldwide pest, long known to be a species complex with more than 40 cryptic – i.e. morphologically indistinguishable – morphotypes (although not all of these are different species). Whiteflies use their mouthparts for sucking up plant sap (phloem) and, like aphids, they can damage plants in a number of ways. They remove plant sap, which reduces the yield and weakens the plants; and they excrete excess phloem sap as honeydew, which provides a substrate for sooty moulds (Perring et al., 2018). Finally, and perhaps most damaging of all, they transmit plant viruses which cause diseases in a wide range of crops. Mixtures of viruses are are spread from plant to plant and field to field by the whitefly vector, B. tabaci.

https://www.flickr.com/photos/iita-media-library/4690366435/in/photolist-89bjdp-88HWNK-6vX388-cEzNrw-6vX3dg-6vX3hv-6w2eyd-6vX3ot-6w2esu-88DNV8-89tmwP-88VdGX-nvXJ3Q-6w2eBS-6w2eH1-nf8Bev-nwhyHs-nf4kz3
Some highly polyphagous members of the B. tabaci complex have become globally invasive crop pests; perhaps the most economically important agricultural and horticultural pests in the World (Malumph et al., 2017; Liu et al., 2022). In Africa, for example, a pandemic of B. tabaci-transmitted Ipomoviruses – have devistated cassava crops throughout East and Central Africa, posing a risk to food security in these regions (Rey and Vanderschuren, 2017).
Some of the species in the B. tabaci complex – scientists have also called them clades or biotypes – differ markedly in their ability to spread plant viruses; and in terms of their endosymbionts (Dickey et al., 2013). Some of these cryptic Bemisia tabaci species thrive better in greenhouses than out of doors, but the adults (see below) can only be told apart by DNA barcoding. In other words, they look identical, but pose different levels of threat to a commercial vegetable grower. The complex as a whole is considered to be a super-vector, transmitting more than 200 plant viruses worldwide (Kanakala and Ghanim, 2019).
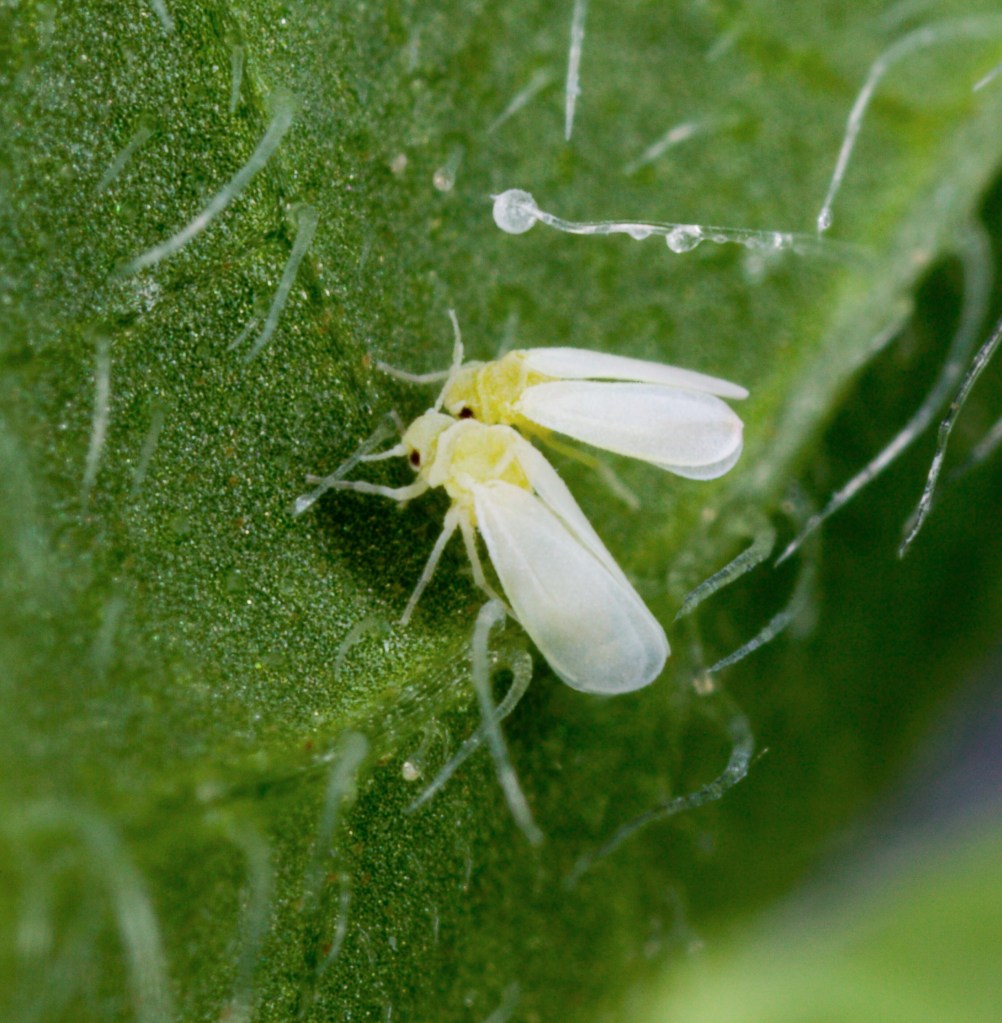

The existence of a complex of biotypes in Bemisia tabaci, with different characteristics, particularly in terms of their imact on commercial crops, has been known for a long time, and resulted in a variety of different names, such as B and Q biotypes. However, these are now considered to be separate species, based on genetic and behavioural data. For example, like all good species, they do not interbreed, i.e. they are reproductively isolated (Sun et al., 2011). Individual males and females do not recognise individuals of the other type/species as being potential mating partners (Wongnikong et al., 2020).
A 2 to 5% divergence in the COI gene sequence, is generally accepted as being indicative of species boundaries in insects (Lee et al., 2017). For example, differences of the order of 3.5 to 4% in the c. 650 base pairs, which make up the genomic sequences in mitochondrial DNA, used for comparisons, indicate the existence of separate species in Bemisia tabaci. Some researchers refer to these different species as mitotypes or mitochondrial haplotypes, because they share similar mtDNA sequences.
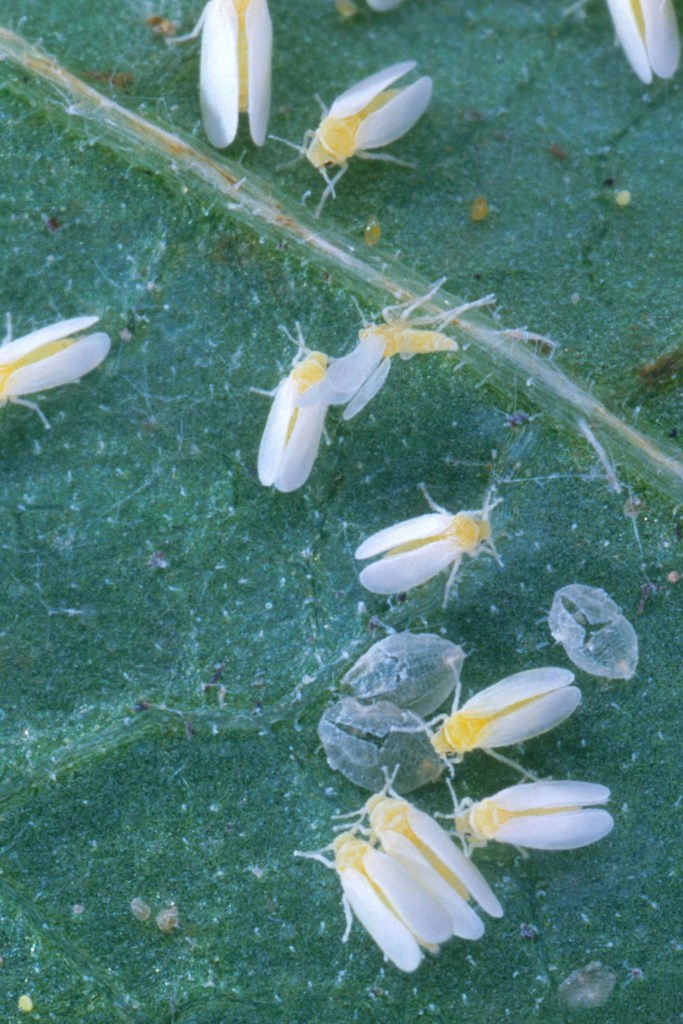

It is generally the case, that species which are reproductively isolated from each other will gradually evolve small changes in their genetic code, for example, at a rate of about 2% per million years (Myr) in the case of mitochondrial DNA (mtDNA), which has a relatively high mutation rate (Herbert et al., 2003). However, not all insect species have such standard nucleotide differences and the extent to which these DNA sequences differ – i.e. percentage terms – between species is somewhat arbitrary.
The rate at which such molecular evolution occurs varies considerably, both between species and within different segments of the genome. Therefore, standard percentage differences in sequence divergence does not always accurately predict species boundaries in insects. In fact, the percentage differences in DNA sequences (both mitochondrial and nuclear) between sister species in a wide range of insects – including termites, bugs, beetles, butterflies, flies and wasps – was found to vary between one and 31% (Cognato, 2006). In other words, DNA data is not an infallible way to delimit species, and if possible, a range of criteria (morphological, geographical, genetic) should be used to define species.
Although it was originally thought that there were no reliable morphological differences between the B. tabaci species – hence the use of the word cryptic – recent research has found that the species which were identified by differences in their DNA, can also be discriminated in terms of their puparial morphology (see below).

Clearly, it is essential to be able to tell different species apart. In the case of pests, in order to identify and control those that present the greatest threat to our food crops; and in the case of beneficial species – most insects are useful in some way or other, supporting and regulating ecosystem services – to preserve species of conservation concern. Putting aside utilitarian arguments, we are, whether we like it or not, custodians of this planet now, and we need to take care of the millions of other species, which have just as much right – more right in fact, because they have been here much longer – than us humans. Knowing who we share the planet with is part of this responsibility. This biodiversity is also of inestimable value; who knows, it may be totally unique in terms of variety in the universe, and each species is a miracle of evolution, containing innovations and adaptations we are only just beginning to understand.
Links
Barcode of life: https://youtu.be/ZImiXgU6bCk
DNA barcoding in zoology and conservation: https://youtu.be/l4Efmpyetoc
Species concepts: https://youtu.be/skCHK_5LrhQ
References
Bossert, S. (2014). Recognition and identification of species in the Bombus lucorum-complex–a review and outlook. bioRxiv, 011379.
Cannon, R. J. C. (2023). Courtship and Mate-finding in Insects: A Comparative Approach. CABI Publishing.
Carolan, J. C., Murray, T. E., Fitzpatrick, U., Crossley, J., Schmidt, H., Cederberg, B., … & Brown, M. J. (2012). Colour patterns do not diagnose species: quantitative evaluation of a DNA barcoded cryptic bumblebee complex. PloS one, 7(1), e29251. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0029251
Chenuil, A., Cahill, A. E., Délémontey, N., Salliant du Luc, E. D., & Fanton, H. (2019). Problems and questions posed by cryptic species. A framework to guide future studies. In From assessing to conserving biodiversity (pp. 77-106). Springer, Cham.
Cognato, A. I. (2006). Standard percent DNA sequence difference for insects does not predict species boundaries. Journal of Economic Entomology, 99(4), 1037-1045.
Dickey, A. M., Osborne, L. S., Shatters Jr, R. G., Hall, P. M., & McKenzie, C. L. (2013). Population genetics of invasive Bemisia tabaci (Hemiptera: Aleyrodidae) cryptic species in the United States based on microsatellite markers. Journal of Economic Entomology, 106(3), 1355-1364.
Hebert, P. D., Cywinska, A., Ball, S. L., & DeWaard, J. R. (2003). Biological identifications through DNA barcodes. Proceedings of the Royal Society of London. Series B: Biological Sciences, 270(1512), 313-321.
Hochkirch, A., Casino, A., Penev, L., Allen, D., Tilley,
L., Georgiev, T., Gospodinov, K. and Barov, B. (2022).
European Red List of Insect Taxonomists. Luxembourg: Publication Office of the European Union.
Hu, P., Lu, L., Hu, S., Da, W., Huang, C. L., Zhang, H., … & Wang, R. (2022). Differentiation of the Chestnut Tiger Butterfly Parantica sita (Lepidoptera: Nymphalidae: Danainae) in China. Frontiers in Ecology and Evolution, 10.
Janzen, D. H., Burns, J. M., Cong, Q., Hallwachs, W., Dapkey, T., Manjunath, R., … & Grishin, N. V. (2017). Nuclear genomes distinguish cryptic species suggested by their DNA barcodes and ecology. Proceedings of the National Academy of Sciences, 114(31), 8313-8318.
Kanakala, S., & Ghanim, M. (2019). Global genetic diversity and geographical distribution of Bemisia tabaci and its bacterial endosymbionts. PLoS One, 14(3), e0213946.
Lee, W., Park, J., Lee, G. S., Lee, S., & Akimoto, S. I. (2013). Taxonomic status of the Bemisia tabaci complex (Hemiptera: Aleyrodidae) and reassessment of the number of its constituent species. PloS one, 8(5), e63817.
Li, X., & Wiens, J. J. (2022). Estimating Global Biodiversity: the Role of Cryptic Insect Species. Systematic Biology.
Liu, Z., Chen, W., Zhang, S., Chen, H., Su, H., Jing, T., & Yang, Y. (2022). Behavioral Responses of Bemisia tabaci Mediterranean Cryptic Species to Three Host Plants and Their Volatiles. Insects, 13(8), 703.
Malumphy C., Eyre D., & Anderson H. (2017). Tobacco, Sweet potato or Silver leaf whitefly. Bemisia tabaci. Plant Pest Factsheet. Department for Environment Food and Rural Affairs. https://planthealthportal.defra.gov.uk/assets/factsheets/Bemisia-tabaci-Defra-Plant-Pest-Factsheet-Feb-2017-2.pdf
Perring, T. M., Stansly, P. A., Liu, T. X., Smith, H. A., & Andreason, S. A. (2018). Whiteflies: biology, ecology, and management. In Sustainable management of arthropod pests of tomato (pp. 73-110). Academic Press.
Ramesh, K. B., Mahendra, C., Kelageri, S. S., Rajna, S., & Subramanian, S. (2022). Distribution and Mitotype Diversity of Bemisia tabaci. Indian Journal of Entomology, 1-5.
Rey, C. & Vanderschuren, H.V. (2017). Cassava mosaic and brown streak diseases: Current perspectives and beyond. Annu. Rev. Virol. 4, 429-452.
Richards, R. A. (2013). The Species Problem: A Conceptual Problem?. The Species Problem-Ongoing Issues, 41. https://www.intechopen.com/chapters/42175
Stork, N. E. (2018). How many species of insects and other terrestrial arthropods are there on Earth. Annual review of entomology, 63(1), 31-45.
Sun, D. B., Xu, J., Luan, J.B. and Liu, S. S. (2011). Reproductive incompatibility between the B and Q biotypes of the whitefly Bemisia tabaci in China: genetic and behavioural evidence. Bull Entomol Res 101:211–220.
Vyskočilová, S., Tay, W. T., van Brunschot, S., Seal, S., & Colvin, J. (2018). An integrative approach to discovering cryptic species within the Bemisia tabaci whitefly species complex. Scientific reports, 8(1), 1-13.
Waters, J. O. E., Darvill, B. E. N., Lye, G. C., & Goulson, D. (2011). Niche differentiation of a cryptic bumblebee complex in the Western Isles of Scotland. Insect Conservation and Diversity, 4(1), 46-52.
Wongnikong, W., van Brunschot, S. L., Hereward, J. P., De Barro, P. J., & Walter, G. H. (2020). Testing mate recognition through reciprocal crosses of two native populations of the whitefly Bemisia tabaci (Gennadius) in Australia. Bulletin of entomological research, 110(3), 328-339.
Zhou, X., Robinson, J. L., Geraci, C. J., Parker, C. R., Flint Jr, O. S., Etnier, D. A., … & Hebert, P. D. (2011). Accelerated construction of a regional DNA-barcode reference library: caddisflies (Trichoptera) in the Great Smoky Mountains National Park. Journal of the North American Benthological Society, 30(1), 131-162.
Very interesting read.
[…] So, it was more a case of the species just not being noticed; or more precisely, not being distinguished from other, very similar bees. It that sense it was cryptic, or hidden. It had presumably existed happily for millions of years, but it only came to our attention relatively recently! Like many so-called cryptic species, see here. […]